Nathan Weinstein
Members-
Posts
13 -
Joined
-
Last visited
Content Type
Profiles
Forums
Downloads
Blogs
Everything posted by Nathan Weinstein
-
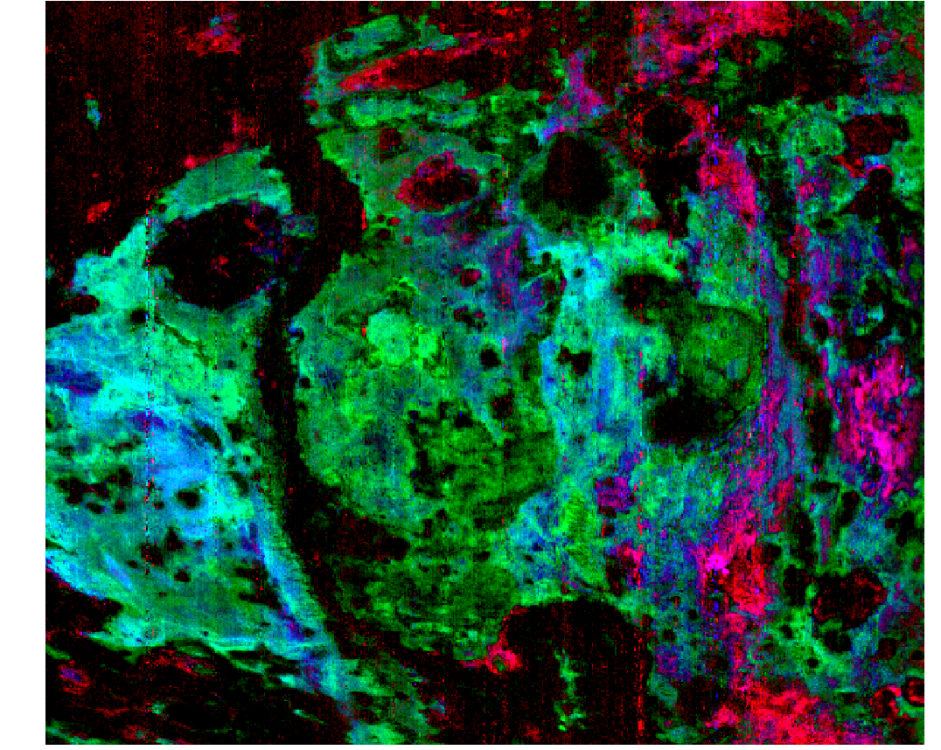
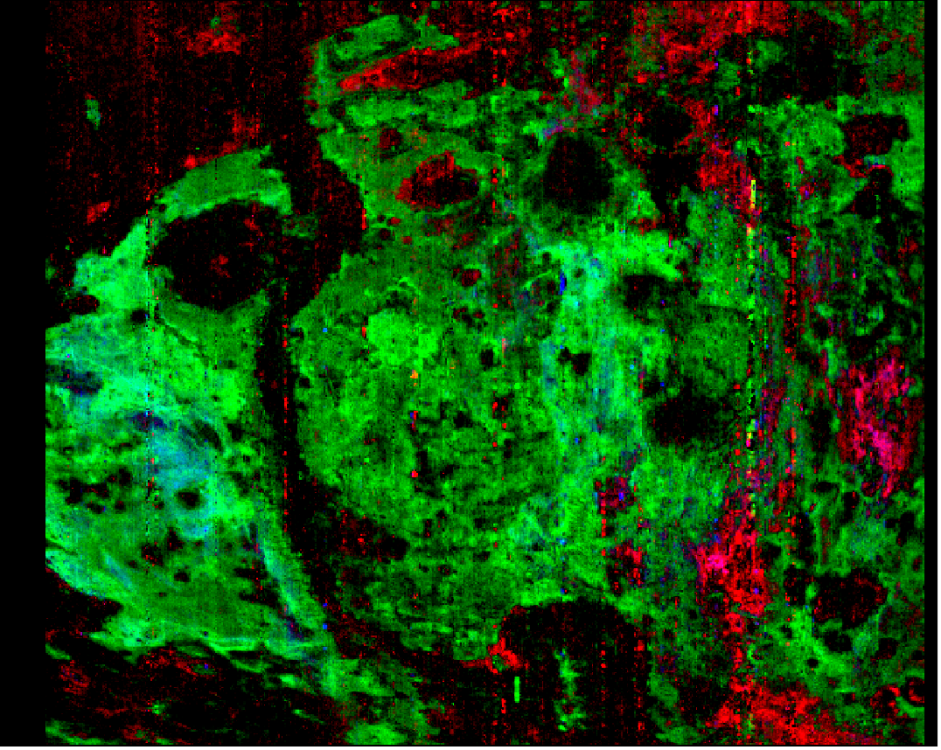
Hello Feng, Thank you for your reply. Yes, of course. As mentioned, I used the FRT00009326 scene. When I load the custom browse product of the pre-calculated summary parameters of SINDEX2, D2300, BD1900_2 (stretch: [0.003, 0.020], [0.010, 0.038], [0.028, 0.058], respectively), by loading from the SU_TER cube – I get this (as should be): But when I am calculating these summary parameters with IF_TER cube by myself with the relevant algebra, I get different values in each parameter matrix (band), which get me this: Thanks, Nathan
-
Hello Ray, Thank you so much for your reply. Attaching my Matlab `browse_product()` code (just change from `.txt` to `.m`) and a spectral library. Please run it and just follow the GUI instructions. I recommend using the default settings for the first time by hitting ‘Enter’ (and clicking on ‘OK’ in stretch parameters choose) through the process. The core function of the summary parameters calculations is inside the main code under the function name `calc_summary_parameters()`, which calls `calc_summary_parameter()` with its nested helper functions for each parameter name. I try to reproduce the work of Wray et al. (2010) with the FRT00009326 scene and bassanite mineral. Also, I’ve uploaded these files to my GitHub repo. Thanks a lot, Nathan Sulfates and Sulfite.txt browse_product.txt
-
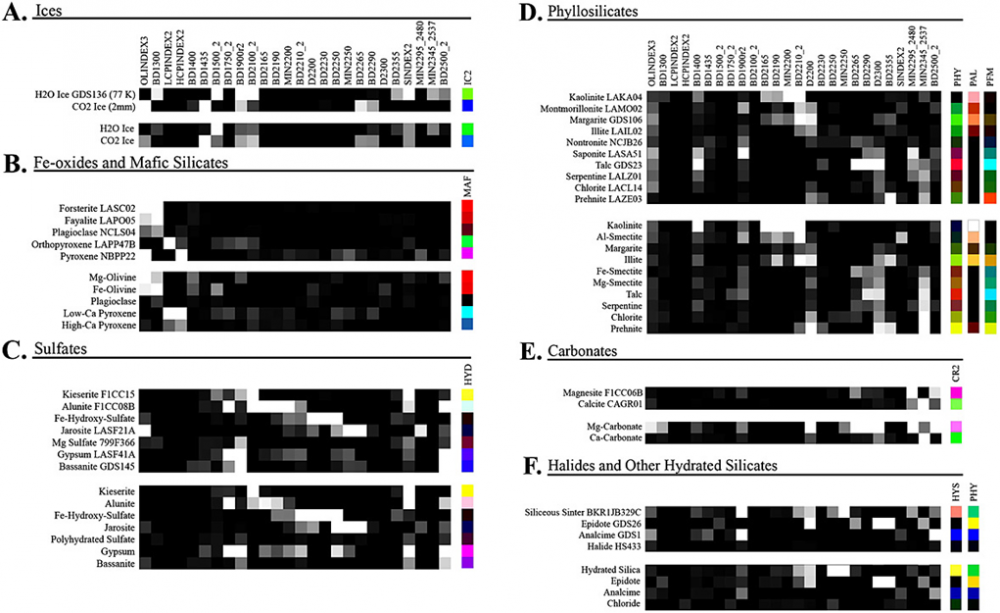
Hello, For my thesis completion: I’m trying to calculate the color of a lab reference mineral with the summary parameters formulations as Viviano et al. (2014) did in this figure (9): For that, I used the relevant summary parameters formulations described in Viviano et al. (2014), table 2, and section 4. But first, to check that my calculations are correctly implemented (using Matlab), I tried to calculate them on an IF_TER cube to reproduce the same browse product as I would get if I opened it by the pre-calculated summary parameters bands in the SU_TER cube (as was shown originally in the data processing pipeline by Seelos et al. 2011). Unfortunately, it wasn’t a matching result. What could be the problem? I appreciate any help you can provide. Nathan
-
hirise Open HiRISE on ENVI 5.6+ together with CRISM
Nathan Weinstein replied to Nathan Weinstein's topic in For data users
Great! Thank you so much! Cheers, Nathan -
hirise Open HiRISE on ENVI 5.6+ together with CRISM
Nathan Weinstein posted a topic in For data users
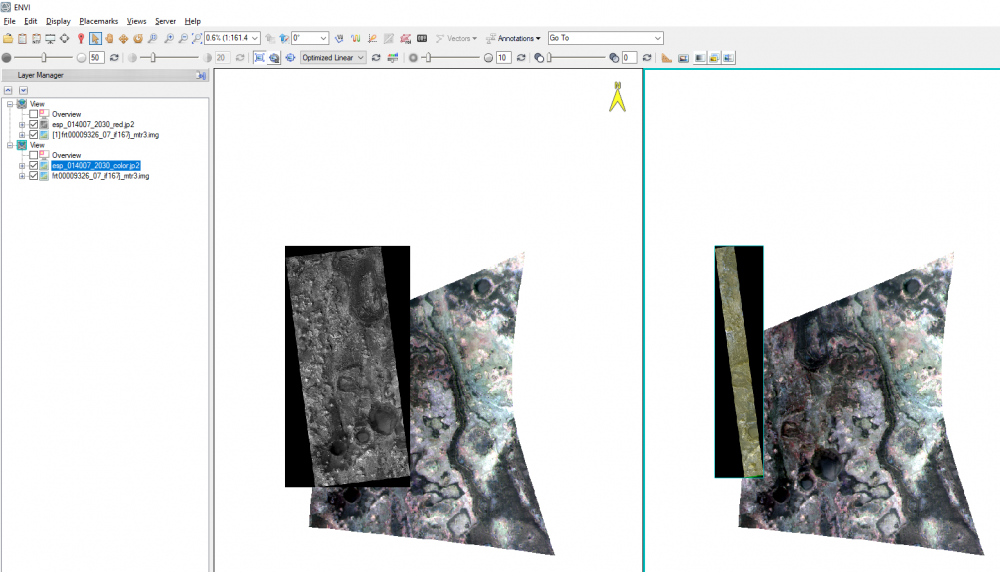
Hello, How to correctly open and overlay map-projected HiRISE and CRISM products? I know that they both should have the same projection (with different pixel size/map scale/spatial resolution). But, unexpectedly they opened from the first pixel's location of the first opened raster layer and not accordingly and independently by the map coordinates of each rasters. See attached. Thanks, Nathan -
I want to calculate the summary product sensitivity matrices of some new analog laboratory material spectrum and represent by a representative browse product RGB as Viviano et. al. (2014) did (attached figure), to use it to search similar RGB corresponding response of CRISM, which also leads me to this question I posted. Thank you, Nathan
- 4 replies
-
- cat
- browse products
-
(and 1 more)
Tagged with:
-
Hi Ray, In the MTRDR pipeline, the BR (8-bit) derived from the SR that is 32-bit. My question is, how it was converted/processed? Thank you, Nathan
- 4 replies
-
- cat
- browse products
-
(and 1 more)
Tagged with:
-
Hello, What do I need to define a new mineral Type Locality? How do I search for new minerals that have not yet been defined with a type locality? I thought of calculating the RGB color of a target mineral lab reference spectrum, with related summary products and then looking for pixels in the related Browse Product scene that have a similar color. Is it a good way? If yes, how do I do the band math on a spectrum instead of on an image? And is there a way to do this with the codes from the directory ‘CAT_ENVI\save_add\CAT_programs\Additions\SummaryParams2014’ ? Thanks, Nathan
-
Hello, How exactly is each summary product converted to an 8-bit RGB channel component of the browsing product? Is there a code procedure in CAT to use with IDL? Are the values in the summary product at a normalized intensity from 0 to 1? If so then is it only left to convert the values to a range [0, 255]? Thanks, Nathan
- 4 replies
-
- cat
- browse products
-
(and 1 more)
Tagged with: